Clonal analysis¶
[1]:
import cospar as cs
[2]:
cs.logging.print_version()
cs.settings.verbosity = 2 # range: 0 (error),1 (warning),2 (info),3 (hint).
cs.settings.set_figure_params(
format="png", figsize=[4, 3.5], dpi=75, fontsize=14, pointsize=3
)
Running cospar 0.2.0 (python 3.8.12) on 2022-01-30 14:45.
[3]:
# Each dataset should have its folder to avoid conflicts.
cs.settings.data_path = "data_cospar"
cs.settings.figure_path = "fig_cospar"
cs.hf.set_up_folders()
Load an existing dataset. (If you have pre-processed data, you can load it with cs.hf.read(file_name).)
[4]:
adata_orig = cs.datasets.hematopoiesis_subsampled()
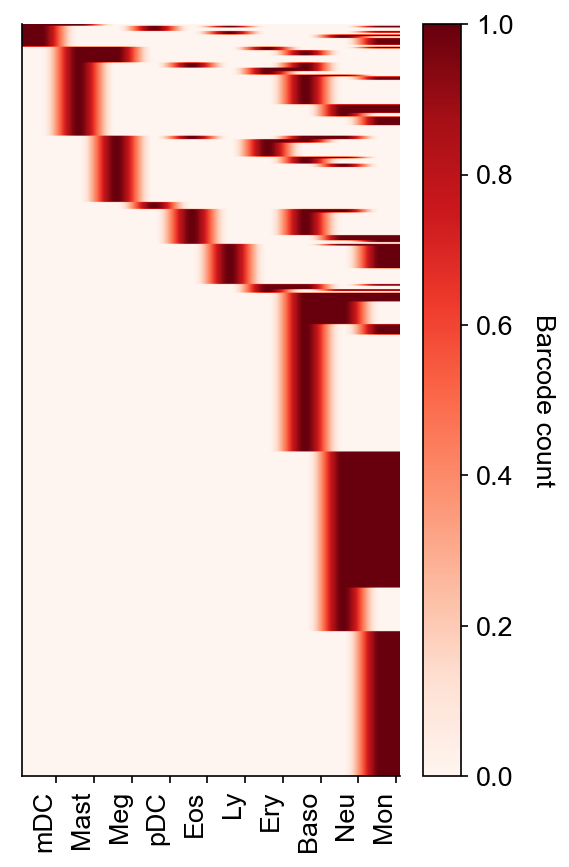
Show barcode heatmap as aggregated into given fate clusters (defined in adata_orig.obs['state_info'])
[5]:
selected_times = None
selected_fates = [
"Ccr7_DC",
"Mast",
"Meg",
"pDC",
"Eos",
"Lymphoid",
"Erythroid",
"Baso",
"Neutrophil",
"Monocyte",
]
celltype_names = ["mDC", "Mast", "Meg", "pDC", "Eos", "Ly", "Ery", "Baso", "Neu", "Mon"]
cs.pl.barcode_heatmap(
adata_orig,
selected_times=selected_times,
selected_fates=selected_fates,
color_bar=True,
rename_fates=celltype_names,
log_transform=False,
binarize=True,
)
Data saved at adata.uns['barcode_heatmap']
[5]:
<AxesSubplot:>
Fate coupling in the underlying clonal data, defined in our package as the normalized barcode covariance between cells annotated in different fates.
[6]:
cs.tl.fate_coupling(
adata_orig, selected_fates=selected_fates, source="X_clone"
) # compute the fate coupling
cs.pl.fate_coupling(adata_orig, source="X_clone") # actually plot the coupling
Results saved as dictionary at adata.uns['fate_coupling_X_clone']
[6]:
<AxesSubplot:title={'center':'source: X_clone'}>

Fate hierarchy constructed from fate coupling of the underlying clonal data, using the neighbor-joining method.
[7]:
cs.tl.fate_hierarchy(
adata_orig, selected_fates=selected_fates, source="X_clone"
) # compute the fate hierarchy
cs.pl.fate_hierarchy(adata_orig, source="X_clone") # actually plot the hierarchy
Results saved as dictionary at adata.uns['fate_hierarchy_X_clone']
/-Lymphoid
/-|
| \-Ccr7_DC
/-|
| | /-Monocyte
| \-|
| \-Neutrophil
/-|
| | /-Baso
| | /-|
| | /-| \-Mast
| | | |
--| \-| \-Eos
| |
| | /-Erythroid
| \-|
| \-Meg
|
\-pDC
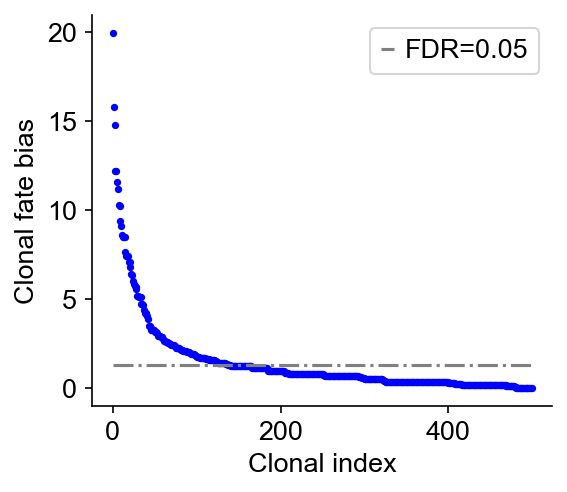
Next, we compute the clonal fate bias, -log(Q-value). We calculated a P-value that that a clone is enriched (or depleted) in a fate, using Fisher-Exact test (accounting for clone size). The P-value is then corrected to give a Q-value by Benjamini-Hochberg procedure. The alternative hypothesis options are: {‘two-sided’,’greater’,’less’}. The default is ‘two-sided’.
[8]:
cs.tl.clonal_fate_bias(
adata_orig, selected_fate="Monocyte", alternative="two-sided"
) # compute the fate hierarchy
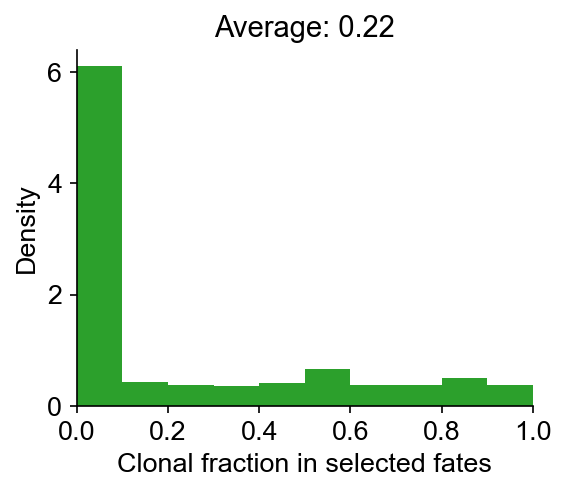
cs.pl.clonal_fate_bias(adata_orig) # actually plot the hierarchy
100%|██████████| 500/500 [00:01<00:00, 442.37it/s]
Data saved at adata.uns['clonal_fate_bias']
[9]:
result = adata_orig.uns["clonal_fate_bias"]
result
[9]:
| Clone_ID | Clone_size | Q_value | Fate_bias | clonal_fraction_in_target_fate | |
|---|---|---|---|---|---|
| 0 | 488 | 58 | 1.037416e-20 | 19.984047 | 0.896552 |
| 1 | 387 | 37 | 1.608291e-16 | 15.793635 | 0.945946 |
| 2 | 227 | 45 | 1.665465e-15 | 14.778465 | 0.866667 |
| 3 | 162 | 42 | 6.210001e-13 | 12.206908 | 0.833333 |
| 4 | 302 | 112 | 6.210001e-13 | 12.206908 | 0.000000 |
| ... | ... | ... | ... | ... | ... |
| 495 | 366 | 9 | 1.000000e+00 | -0.000000 | 0.222222 |
| 496 | 46 | 18 | 1.000000e+00 | -0.000000 | 0.222222 |
| 497 | 408 | 27 | 1.000000e+00 | -0.000000 | 0.259259 |
| 498 | 6 | 4 | 1.000000e+00 | -0.000000 | 0.250000 |
| 499 | 463 | 3 | 1.000000e+00 | -0.000000 | 0.333333 |
500 rows × 5 columns
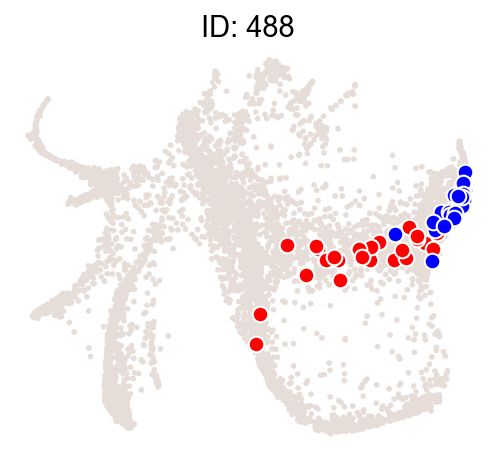
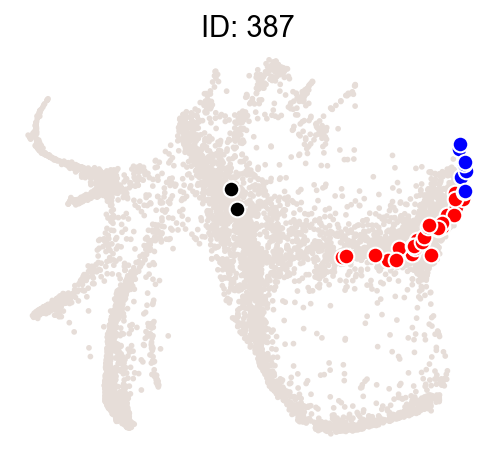
Illustrate some most biased clones.
[14]:
ids = result["Clone_ID"][:2]
cs.pl.clones_on_manifold(
adata_orig,
selected_clone_list=ids,
color_list=["black", "red", "blue"],
clone_markersize=15,
)